Part Of: Biology sequence
Content Summary: 3000 words, 15 min read
The Basic Facts of Abiogenesis
From the geochemistry of the early Earth, life emerged. The abiogenesis phenomenon is interesting in at least two contexts:
First, it is one of the most difficult unsolved scientific problems in the 21st century. Common descent describes how all of the marvelous diversity of biology stem from a single source: a single celled organism called the Last Universal Common Ancestor (LUCA). We have a clear mechanistic understanding for how this complexification was possible: evolution by natural selection.
But natural selection relies on the machinery of genetic inheritance to produce complexity. How was it possible to create the sophisticated nanomachinery of LUCA (e.g., ATP synthase) before natural selection took effect? What other process besides natural selection can produce complexification?

Second, abiogenesis research is relevant to questions in astrobiology:
Could extraterrestrial life have begun much earlier?
- Gen 1 stars (born 13.5-13 Ga) had no exoplanets because the heavy elements required for planetary cores didn’t exist (supernova nucleosynthesis arrived later).
- Gen 2 stars (born 13-10 Ga) may have been able to support life, but these exoplanets were small, terrestrial, and less metallic – unable to furnish geomagnetism nor plate tectonics.
- Gen 3 stars (born 10-0 Ga) can support life. Exoplanets in this era now include gas giants, and are highly metallic. Geomagnetism protects us from UV radiation, and plate tectonics promote mineral diversity.
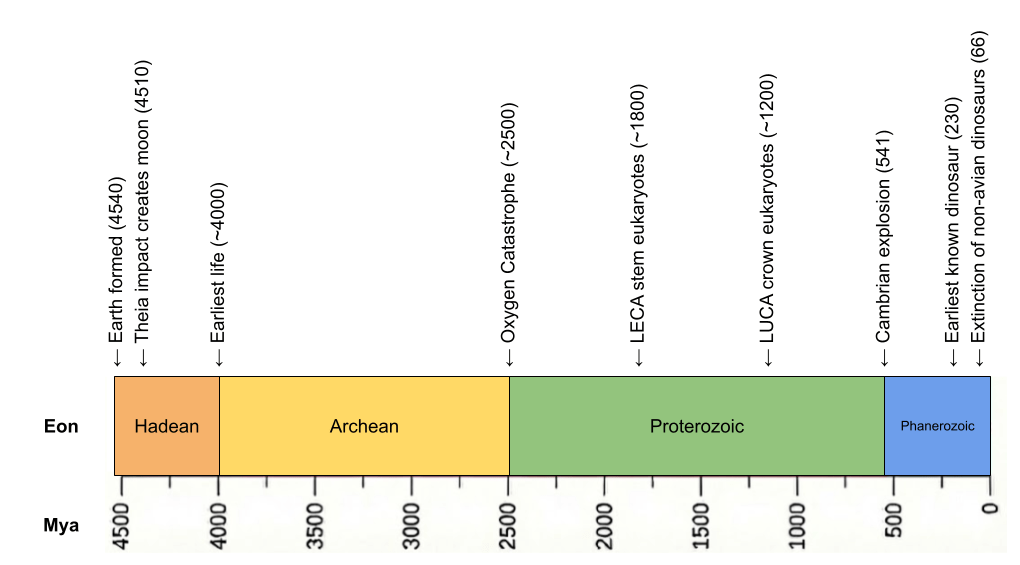
Is abiogenesis easy or hard? The Earth was born 4.6 Ga, with the Theia impactor at 4.54 Ga creating the moon and the Moneta impactor at 4.51 Ga providing siderophilic veneer. Despite such impacts, zircon evidence now suggests a cool early earth, with oceans appearing as early as 4.4 Ga. Critically, our evidence for life arrives very early (Isua banded-iron formations at 3.8 Ga, enriched carbon-12 in zircon graphite at 4.1 Ga). Abiogenesis occurred only ~500 million years after the sterilizing impact of Moneta! Speed is evidence of ease.
If the universe has billions of potentially habitable planets, why haven’t we found any aliens? One explanation to the Fermi Paradox is the Great Filter – some incredibly difficult barrier that prevents life from becoming spacefaring. The scary part is we don’t know where this filter is: behind us (we got incredibly lucky to make it this far) or ahead of us (something typically destroys civilizations before they can spread across the galaxy). If abiogenesis is indeed easy, this might mean the Great Filter is ahead of us.
LUCA from Paleobiogeography
To understand the journey of abiogenesis, we must first understand the destination. What do we know about LUCA?
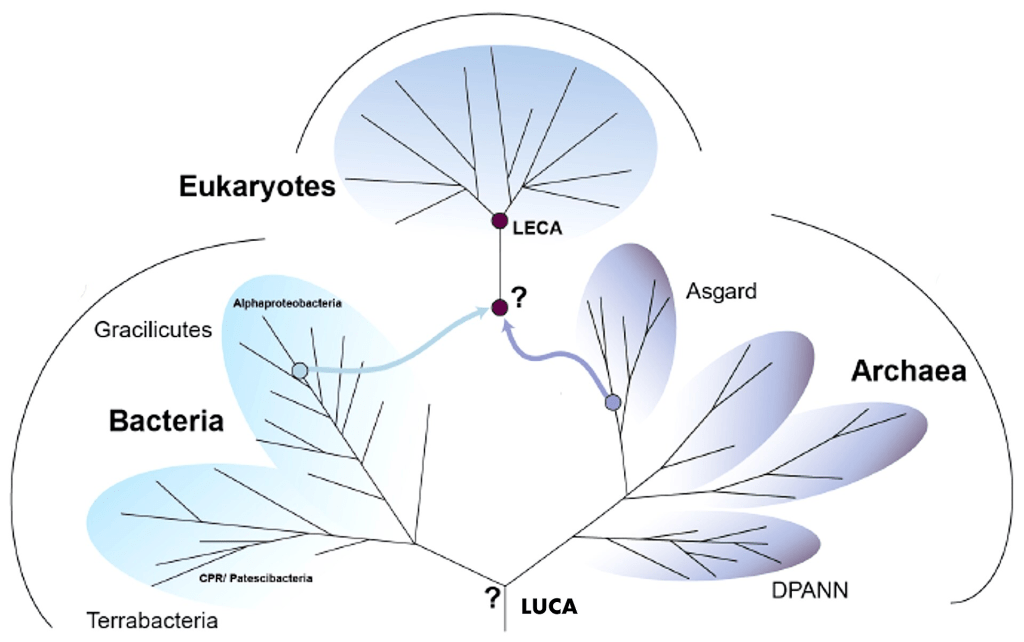
The history of life constrains our search. Two domains of simple life (prokaryotes) emerged in the Hadean: bacteria and archaea. But eukaryotes (complex life) emerged much later, as a result of endosymbiosis between the two families.
Life was microbial for the first 80% of Earth’s history. But in 1.0 Ga, multicellularity was invented. Starting at that time, multicellularity was invented multiple times, but it only really took off in crown group eukaryotes.
LUCA must have been anaerobic. For the first half of Earth’s history, oxygen was locked in water and rocks – there was no free oxygen in the atmosphere. Only when photosynthesis caused the Oxygen Catastrophe in 2.4 Ga did atmospheric oxygen go from 0 to 10%.
LUCA is simple (prokaryotic), microbial (unicellular), and anaerobic.
LUCA from Phylogenetics
We can learn more about LUCA from genetic analyses.
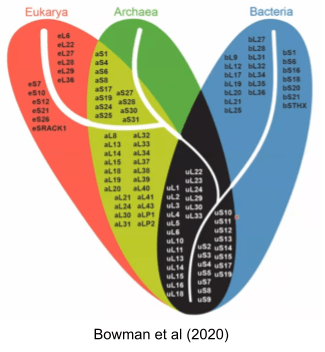
Genes conserved across all species constitute what is called the Universal Gene Set of Life (UGSL), and consists of less than 100 genes (Harris et al 2003). Not surprisingly, the UGSL is dominated by translation-related genes. Here are those ribosomal genes in black:
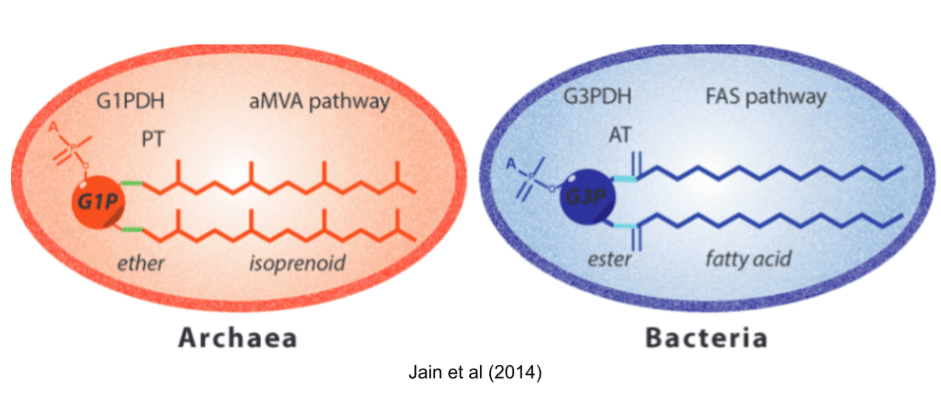
Many differences between archaea and bacteria arose after they diverged. But some differences are more troubling. First among these is the Lipid Divide. Both domains build cells with the phospholipid bilayer membranes, so it is natural to suspect that LUCA had a similar coat. But almost all biochemical details are mirrored! They use different glycerol backbones, different hydrophobic chains (isoprenoid vs fatty acids), different links to those chains (ether vs ester), and different biosynthetic pathways (FAS+AT vs aMVA+PT).
There are three major theories for the lipid composition of LUCA:
- Heterochiral theories. LUCA had both lipid biochemistries available. Heterochiral membranes were recently found to be viable (Caforio et al 2018).
- Thermoreduction theories. Many archaea are adapted to extreme environments. So LUCA had a bacterial phospholipid, and the more robust archaeal lipids were derived to support more extreme niches.
- Protocompartment theories. LUCA didn’t use phospholipids at all! It used a simple fatty acid membrane, or coacervate droplets, or mineral pores to achieve compartmentalization.
Another divergence is even more troubling: bacteria and archaea use completely different DNA replicase proteins (Forterre et al 2013).
Neither divide has a satisfying resolution. Each of the three lipid hypotheses faces serious objections, and the DNA replicase situation is arguably worse: there is no consensus account for how two non-homologous replication machineries could descend from a single ancestor, which has led some (e.g., Forterre 2006) to argue that DNA itself was a viral invention acquired independently by the bacterial and archaeal lineages. Any portrait of LUCA that papers over these divides is overconfident.
LUCA from Biochemistry
At the atomic level, organisms are built with CHNOPS: Carbon, Hydrogen, Nitrogen, Oxygen, Phosphorous, and Sulfur.
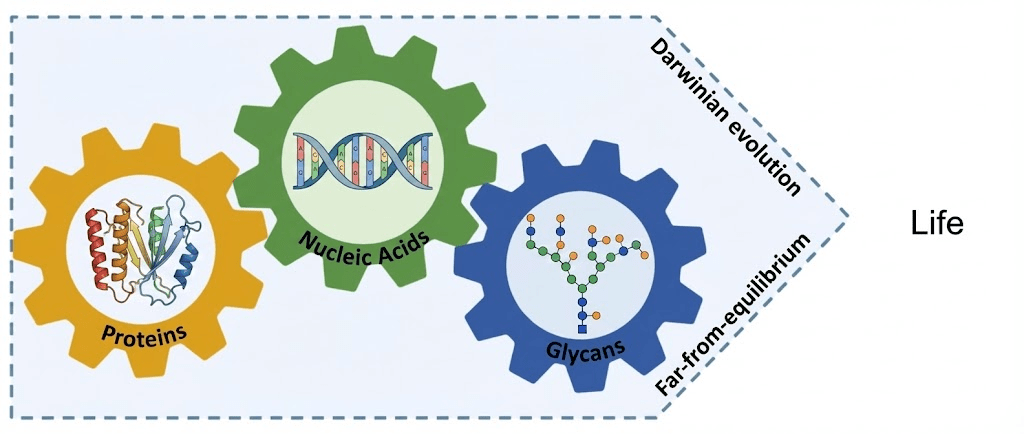
At the molecular level, life is composed of three biopolymers.
- Glycans are made out of monosaccharides (e.g., glucose).
- Proteins are made out of amino acids.
- Nucleic acids are made out of nucleotides.
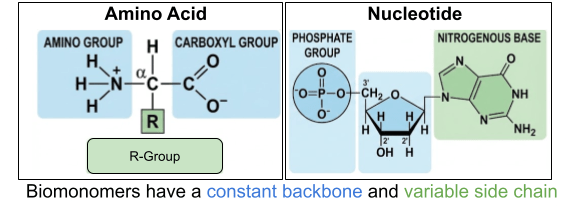
Most of our discussion will center on proteins and nucleic acids. Their constituents are themselves large, complex molecules. These monomers are built on an unchanging backbone and a variable side chain. For nucleotides, there are 4 available side chains (purine and pyrimidine nitrogenous bases). For amino acids, there are 20 available side chains (nonpolar, polar-uncharged, polar-positive, polar-negative R groups).
Organic polymers like polyesters aren’t particularly flexible. But biopolymers manifest extreme functional sophistication. And the polyfunctionality of nucleic acids and glycans must not be understated (Matange et al 2025).
Biopolymers also exhibit functional interdependence. Condensation of nucleotides is catalyzed by proteins, and condensation of amino acids is catalyzed by RNA. Biopolymers are heterocomplementary: proteins can recognize and bind to proteins, DNA, polyglycans, and small molecules. Nonbiological organic polymers do not manifest heterocomplementarity.
LUCA from Microbiology
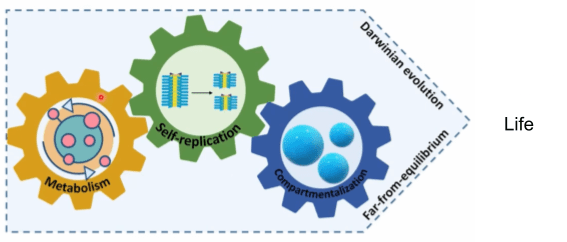
Another way to approach LUCA is to catalog biological universals: mechanisms shared by all life forms in existence. These universals can be organized in three categories: self-replication, metabolism, and compartmentalization. Let’s discuss each in turn.
The central genius of life was the discovery of proteins. By chaining amino acids together in long biopolymers, these polypeptides fold into arbitrary shapes. This invention is so powerful because in chemistry, structure determines function: arbitrary shapes unlock myriad potential behaviors. Proteins can form fibers, motors, containers, transporters, sensors, signals, optical devices, adhesives, pores, brushes, and pumps.
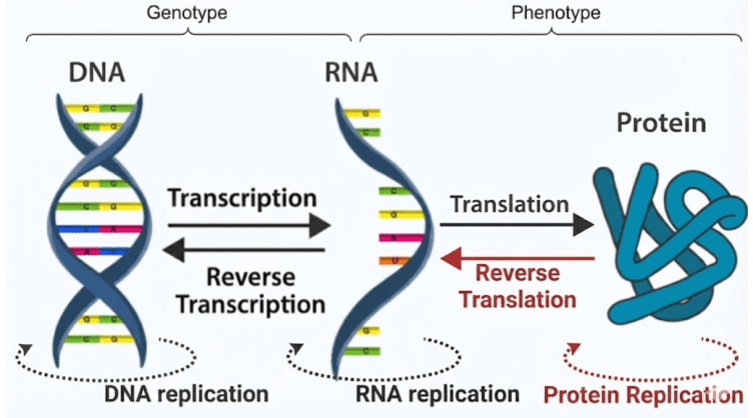
Proteins aren’t synthesized randomly – they are produced by template. Nucleotides combine together in long biopolymers known as nucleic acids. Double-stranded DNA is copied into single-stranded mRNA via the process of transcription. Then, during protein synthesis in the ribosome, codons (RNA triplets) specify which amino acids to attach in which order. This process of translation is defined by a codon-amino acid map known as the genetic code.
A gene is simply a patch of RNA that completely specifies a particular protein sequence. Your genome (genotype) is nothing more than a recipe for a proteome (phenotype). When philosophers speak of an innate desire to survive, that motivation must be carried by proteins. Natural selection operates on protein design. Nothing more and nothing less.
The Central Dogma describes the flow of information in biopolymers. The black arrows are allowed processes; the red arrows are not observed. Proteins cannot replicate because they are incapable of base pairing. Once information gets into a protein, it cannot get out again!
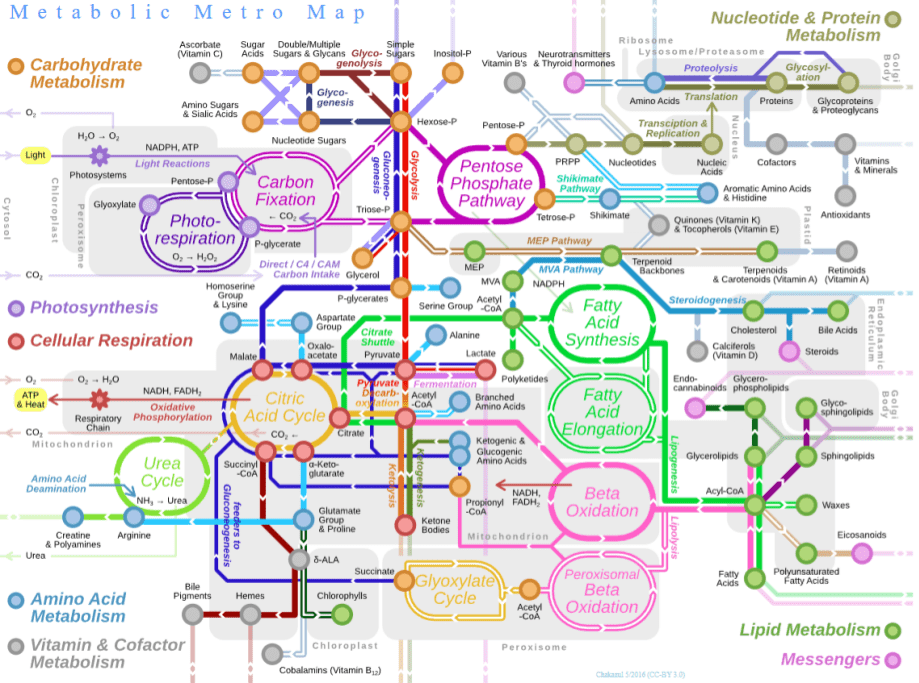
Why is life so fixated on proteins? To better understand this, it helps to consider metabolism. This is a chemical reaction network which provides two basic functions:
- Catabolism (“energy metabolism”), a destructive process which breaks down complex compounds. This prominently includes pathways of cellular respiration, a controlled version of combustion in which glucose is slowly converted into carbon dioxide, water, and energy.
- Anabolism (“carbon metabolism”), a constructive process which biosynthesizes new organic compounds. This includes pathways of carbon fixation (like Acetyl-CoA), which converts simple inorganic carbon dioxide into complex organics.
Enzymes are protein-based catalysts that reduce the activation energy of thermodynamically favorable (exergonic) reactions, speeding them up by many orders of magnitude. Some metabolic steps, however, are thermodynamically disfavorable (endergonic) and will not proceed spontaneously. Enzymes handle these by coupling them to a favorable reaction, making the combined process exergonic. The enzyme then catalyzes that coupled reaction in the usual way — by lowering its activation energy.
The 20 amino acids have a limited chemical repertoire; cofactors extend it — enabling electron transfer, redox chemistry, and group transfers that amino acid side chains cannot perform alone. There are two categories of cofactors: metal ions (e.g., Mg²⁺, and Fe²⁺), and organic molecules (i.e., coenzymes), many of which are derived from B vitamins (e.g., NAD). Without its cofactor, an enzyme (called an apoenzyme) is typically inactive. The complete, functional assembly is the holoenzyme.
Finally, while the Central Dogma traditionally depicts a linear flow of information from DNA to RNA to Protein, metabolism reveals that this dogma is actually a loop. Proteins are not just passive end-products; they are the active machinery required to synthesize and maintain the very DNA and RNA that encode them, creating a self-sustaining cycle of synthesis and regulation.
Modern metabolism and self-replication does not occur in an undifferentiated “primordial soup”. They occur inside impermeable membranes made of phospholipids.
Compartmentalization does a lot of heavy lifting at once: it shields nucleic acids from genetic parasites, buffers the system against environmental stress, and locally concentrates enzymes and substrates to speed reactions. By keeping metabolites and genetic material from simply diffusing away, it preserves hard-won products and information. Selective, protein-based gates regulate traffic—letting in foodstuffs, exporting waste—and can couple chemiosmosis to maintain a strong proton-motive force (≈3 pH units and ~200 mV), effectively a “cellular battery” that powers metabolic work. Finally, a stable compartment provides the physical unit that can reliably copy itself and propagate success via binary fission.
LUCA from Virology
Previous sections reconstructed LUCA as a cell. But modern oceans contain ten virions for every microbe. Viruses coevolve with cells, outnumber them, and may even predate them. Any reconstruction of early life that ignores them is incomplete.
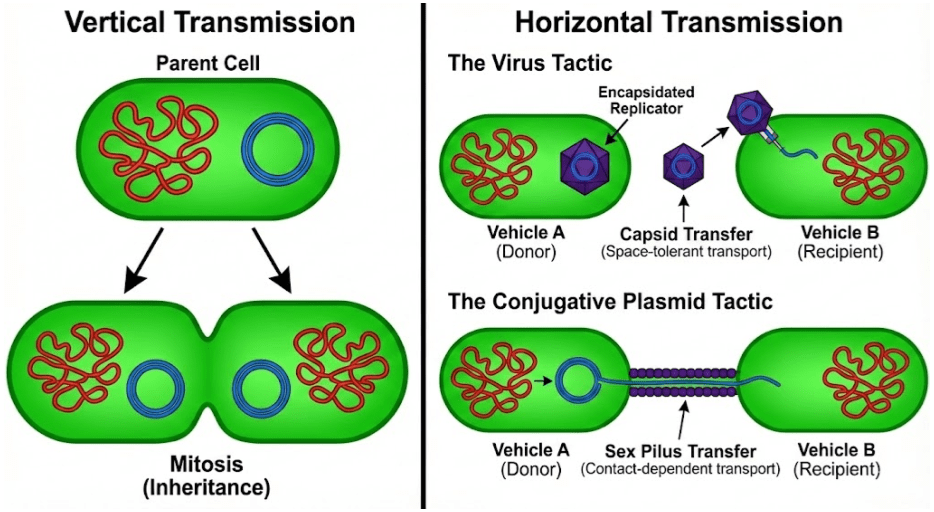
Once inside the cell, virulent viruses initiate the lytic cycle. Their genes hijack the host cell’s machinery to make more copies of themselves, ultimately rupturing the membrane and spilling newly-minted virions into the environment. They rely on horizontal transmission, in contrast to chromosomes which rely on vertical transmission via mitosis.
Viruses are not the only mobile genetic element (MGE) using horizontal transmission. Cells can contain small (often circular) nucleic acid outside of the chromosomes. These plasmids don’t kill the cell to transfer their DNA, but instead use conjugative sex pili to copy themselves into neighboring cells. Plasmid genetic material can be more similar to viruses than the chromosome. Some plasmids and viruses differ by no more than a capsid gene (Krupovic & Bamford 2010).
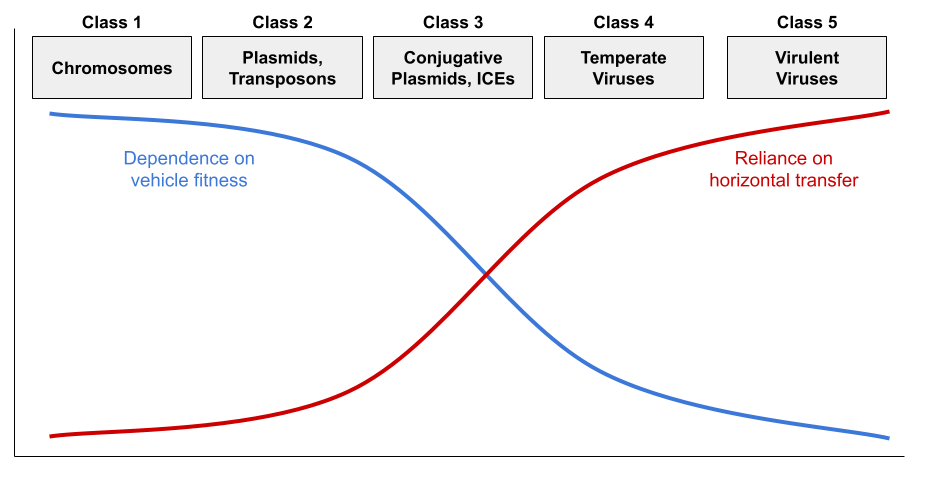
But virulence and conjugation are extremes. While virulent viruses are indifferent to cellular fitness, temperate viruses benefit from vertical transmission via the lysogenic cycle. They are thus less pathogenic. Some plasmids cannot construct the sex pilus and are capable of horizontal transmission only rarely. Jalasvuori (2012) shows that nucleic acids can be organized into a continuum:
This framework explains the distribution of replicator properties:
- Class 3-5 all sense damage to the host cell, and initiate horizontal transfer when they detect vehicle stress. For example, herpesviruses reactivate from latency under host stress, producing cold sores.
- Class 1 retains the core genome (e.g., DNA replication). Class 2-3 retain the accessory genome (e.g., virulence, antibiotic resistance).
- Class 1-5 all retain addiction modules (a.k.a., MGE stabilization modules) like the toxin-antitoxin (TA) system (Mendoza-Guido & Rojas-Jimenez 2025). Many plasmids produce both a toxin and the antitoxin. Since the toxin has a longer half-life, loss of the plasmid entails death of the host. The restriction-modification (RM) system works in a very similar way (Kobayashi 2001).
There are three hypotheses for the origin of viruses:
- The regression hypothesis claims that viruses are cells that progressively lost genetic information in their journey towards obligate parasitism.
- The escape hypothesis claims that viruses originate from MGEs that gained increasingly sophisticated methods of horizontal transfer.
- The virus-first hypothesis claims that viruses predate LUCA, originating from the primordial pool of replicators prior to the evolution of cells.
Dependent niches incentivize genome reduction. Mitochondria have transferred many genes to their host cells’ nuclei (Andersson & Kurland 1998), and parasitic bacteria like Rickettsia have shed much of theirs (Diop et al 2019). As obligate parasites, viruses should face the same reductive pressure — but what if they weren’t always parasitic? The discovery of giant viruses (in the phylum Nucleocytoviricota), with the first known translation-related machinery in a virus, was a revelation. In the spirit of the regression hypothesis, Boyer et al (2010) described them as living fossils of this fourth domain. But, while the relationship between reduction and parasitism is strong, recent phylogenetic work (Monttinen et al 2021) does not support this hypothesis.
The escape hypothesis accounts for some viral genes – particularly those encoding capsid proteins – which have clear cellular homologs. But viral hallmark genes, especially those involved in replication, are shared across diverse viral lineages yet absent from cellular genomes (Koonin et al 2006). If these genes escaped from cells, their cellular homologs should exist. Krupovic et al (2019) propose a chimeric origin that reconciles the escape and virus-first views. They argue that structural genes were captured from hosts, but replicator genes descend from primordial self-replicating elements that predate cells.
Theoretical models reinforce this view — selfish genetic elements arise inevitably in any replicator system, suggesting that genetic parasites have been components of life from the very beginning. Before replicators evolved error correction mechanisms, early replicators also faced a hard physical constraint known as Eigen’s error threshold: RNA replication error rates impose an upper bound on genome size. Viroids, the simplest known replicators at ~300 nucleotides, sit near this boundary. If the chimeric model is correct and viral replicator genes descend from primordial self-replicating elements, viroids may be the closest living relatives of those elements. Their structural simplicity, their lack of protein-coding capacity, and the recent discovery of viroid-like circular RNAs across diverse environments (Lee et al 2023) are all consistent with deep ancestry.
Looking Beyond the Root
The portrait above shows LUCA as anaerobic, prokaryotic, protein-dependent, chromosomally organized, besieged by viruses. But every feature of that portrait is also a choice made from a menu of alternatives that were never taken:
- Alternative codes. Of ~500 amino acid species, only 20 are included in the genetic code, even though codons could distinguish up to 64. Central metabolites like ornithine, GABA, and β-alanine are biosynthesized but never translated.
- Alternative replicator backbones. 5-carbon sugars (ribose, deoxyribose) are a strange choice for a nucleic acid backbone (Eschenmoser 1999). Chemists have built alternative replicators from 4-carbon sugars (TNA) and 3-carbon sugars (GNA).
- Alternative protein backbones. Proteins are built from α-amino acids, but foldamers made from β, γ, and δ-amino acids can be more resistant to proteolysis and heat denaturation than their α-counterparts (Gellman 1998).
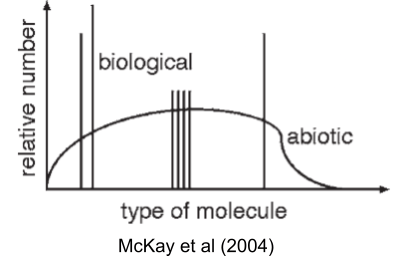
McKay (2004) captures this puzzle visually: nonbiological chemistry produces smooth distributions of organic molecules, but life uses only a sparse set of spikes against that background.
Each of these alternatives works better than what life uses, on some axis. So the interesting question isn’t whether LUCA could have been different; it’s why it wasn’t. Why haven’t we found shadow biospheres running on TNA, or β-peptide proteomes, or 34-amino-acid codes? Two possibilities: either we haven’t looked hard enough, or these choices were frozen so early and so deeply that no later lineage could back out of them. Both answers require understanding what happened before LUCA: the evolution of the ribosome, the freezing of the code, the crystallization of a chromosome out of competing replicators.
Beyond the root of the tree of life lies the origin. That’s the subject of our next post.
References
- Andersson & Kurland (1998). Reductive evolution of resident genomes
- Bowman et al (2020). Root of the Tree: The Significance, Evolution, and Origins of the Ribosome
- Boyer et al (2010). Phylogenetic and Phyletic Studies of Informational Genes in Genomes Highlight Existence of a 4th Domain of Life Including Giant Viruses
- Caforio, A., et al. (2018). “Converting Escherichia coli into an archaebacterium with a hybrid heterochiral membrane.”
- Diop et al (2019). Paradoxical evolution of rickettsial genomes
- Eschenmoser (1999). Chemical etiology of nucleic acid structure.
- Forterre (2006). The origin of viruses and their possible roles in major evolutionary transitions.
- Forterre et al (2013) Origin and Evolution of DNA and DNA Replication Machineries
- Gellman (1998). Foldamers: A manifesto.
- Harris et al (2003). The genetic core of the universal ancestor.
- Jain et al (2014). Biosynthesis of archaeal membrane ether lipids
- Jalasvuori (2012). Vehicles, replicators, and intercellular movement of genetic information: evolutionary dissection of a bacterial cell.
- Koonin et al (2006). The ancient virus world and evolution of cells.
- Kobayashi (2001). Behavior of restriction–modification systems as selfish mobile elements and their impact on genome evolution.
- Krupovic et al (2019). Origin of viruses: primordial replicators recruiting capsids from hosts.
- Krupovic & Bamford (2010). Order to the viral universe
- Lee et al (2023). Mining metatranscriptomes reveals a vast world of viroid-like circular RNA
- Matange et al (2025). Biological polymers: evolution, function, and significance.
- McKay (2004). What Is Life—and How Do We Search for It in Other Worlds?
- Mendoza-Guido & Rojas-Jimenez (2025). Beyond plasmid addiction: the role of toxin-antitoxin systems in the selfish behavior of mobile genetic elements.
- Monttinen et al (2021). The genomes of nucleocytoplasmic large DNA viruses: viral evolution writ large