Part Of: Demystifying Life sequence
Content Summary: 1200 words, 12 min read
Central Thesis Of Molecular Biology
In every cell of your body, there exist molecules called deoxyribonucleic acid. Such cells come in four flavors and (due to their atomic shape) tend to pair up and create long strings. These strings become very long, over two inches when held end-to-end (but of course, they fold up dramatically so each can comfortably inhabit a single cell). Since your cells have about 46 inches worth (six billion molecules), each cell contains twenty-three unique strings. They look like this:
Let us refer to these strings as chromosomes, and to all of them collectively as the human genome. Finally, since typing “deoxyribonucleic acid” is fairly onerous, we will use the acronym DNA.
In 1956, Francis Crick presented his Central Thesis Of Molecular Biology, which describes how the causal chain DNA → RNA → amino acids → protein ultimately motivates every trait of every living organism. A gene is a sequence of DNA that encodes a protein. A genotype (some animal’s unique DNA) explains phenotype (that animal’s unique traits). Genotype-phenotype maps (GP-maps) turn out to be very important in what follows.
Duplication vs. Mutation
Every time a cell duplicates itself (mitosis), its DNA is copied into the new cell. If every cell contains exactly the same code, how can they be different? The basic explanation of cellular differentiation involves feedback loops in the genetic causal chain (collectively named the Gene Regulatory Network). When a lung cell is duplicated, for example, it inherits not just the entire genome, but also proteins for activating lung genes and deactivating other code.
Germ cells are created by a different process entirely. Instead of genome duplication (mitosis), germ cells inherit what is essentially half a genome, in a process known as meiosis. Here’s how these two processes work:
Recall that deoxyribonucleic acid is a collection of atoms. Replicating such a fragile object is imperfect. There are many kinds of ways the process can go wrong; for example:
- Replacement Mutation (e.g., AGTC → AATC)
- Duplication Mutation (e.g., AGTC → AGGTC)
- Insertion Mutation (e.g., AGTC → AGATC)
How many mutations do you have? While you can always get your DNA sequenced to find out, the answer for most people is about sixty.
The Landscape Of Gene-Space
Consider all animals whose genome is three molecules long. How many genetically unique kinds of these animals are there? Recall there are four kinds of DNA: cytosine (C), guanine (G), adenine (A), or thymine (T). We can use the following formula:
Here we have 3^4 = 81 possible genotypes in this particular gene-space. To visualize this, imagine a 4-sided Rubik’s Cube: each dimension is a slot, each cube a particular genotype in the space.
But humans have approximately three billion base pairs; the size of a realistic gene-space is almost incomprehensibly large (4^3,000,000,000), far exceeding the number of atoms in the universe. Reasoning about 3D cubes is easy, reasoning about 3,000,000,000-D hypercubes is a bit harder. So we employ dimension reduction to aid comprehension. If you laid all 4^3,000,000,000 numbers out on a two dimensional matrix, each cell would be so tiny that the surface would appear continuous. We have arrived at our first metaphor identification:
- A Genotype Is A Location
We can summarize our discussion of mitosis, meiosis, and mutation as follows:
- An Organism Is A Stationary Point
- Birth Is Point Creation, Death Is Point Erasure.
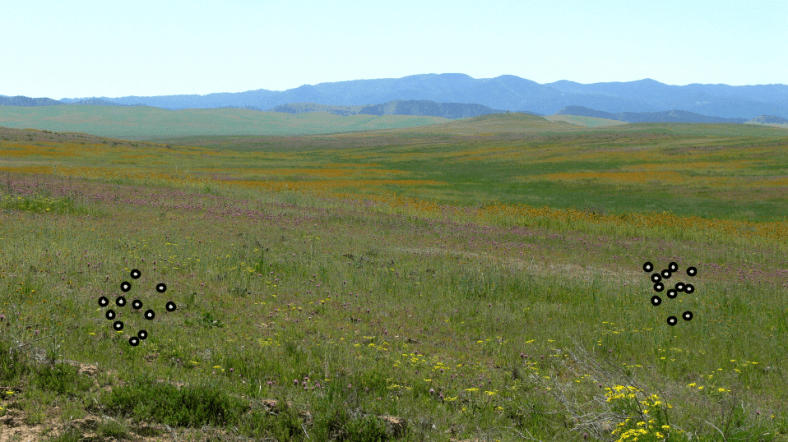
Finally, let us explore the concept of genetic distance. From our toy gene-space, let me take seven nodes and draw lines indicating valid replacement mutations between them.
The key observation is that distances vary. Many nodes are connected via one mutation, but the minimum distance from top (ATG) to bottom (CCC) is three mutations. In other words:
- Varying Genome Differences Are Varying Distances
Our gene-space landscape, then, looks something like this:
Species Are Clusters
What is a species? After all, there is no encoding of the word “jaguar” in the jaguar genome. Rather, members of a species share more genetic similarities to one another than other organisms. In terms of our metaphor:
- A Species Is A Cluster Of Points
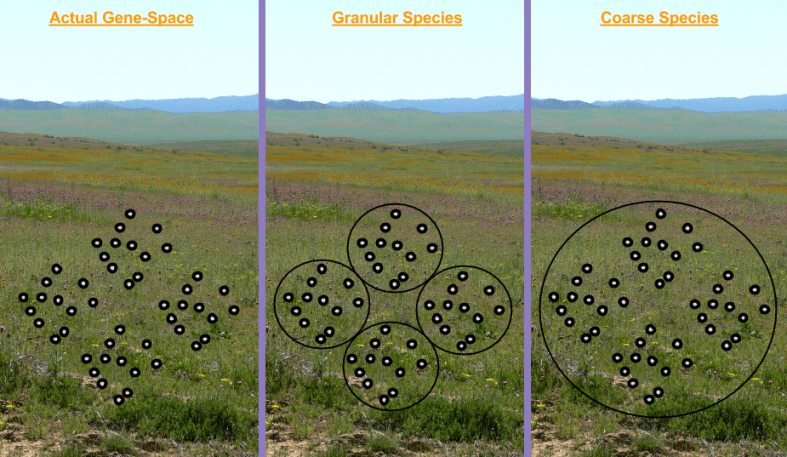
In the above landscape, we might have two species. But there are many ways to cluster data. Consider these competing definitions:
Which clustering approach is correct? It depends on the scale of our axes:
- If we chose Granular but are too “zoomed in”, we have accidentally defined four new species of Shih Tzu.
- If we chose Course but are too “zoomed out”, we have accidentally defined Mammal as its own species.
The point is that scale matters, and we should define species on a scale that makes good biological sense. The most popular scale is that defined by successful interbreeding (i.e., produce fertile offspring). For greater distances (large genetic dissimilarity), such interbreeding is impossible. We therefore constrain the size of our specie clusters by maximum interbreeding distance.
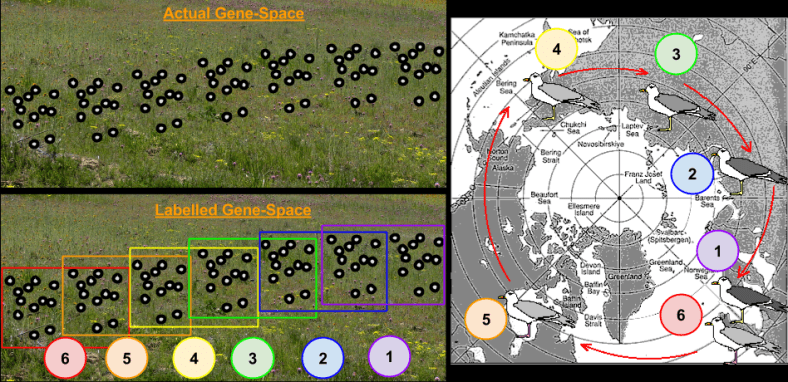
The approach just outlined is the one in use today. However, any man-made criteria for categorizing reality has its stretch points. For example, consider ring species.
Consider the Larus gulls’ populations in the above image. These gulls habitats form a ring around the North Pole, not normally crossed by individual gulls. The European herring gull {6} can hybridize with the American herring gull {5}, which can hybridize with the East Siberian herring gull {4} which can hybridize with Heuglin’s gull {3}, which can hybridize with the Siberian lesser black-backed gull {2}, which can hybridize with the lesser black-backed gulls {1}. However, the lesser black-backed gulls {1} and herring gulls {6} are sufficiently different that they do not normally hybridize.
Genetic Drift Is Random Travel
Landscapes without movement aren’t very interesting. With our brand-new concept as Species As Clusters, let’s see if we can make sense of travel.
Consider the phenomenon of population bottleneck. Many factors may contribute to population reduction (e.g., novel predators). Often, the survivors are just lucky. Descendants of the survivors tend to be more similar to them than the average genome of the original species. By this process, bottlenecks induces change in the species as a whole:

Why wouldn’t such movement cancel itself out in the long run? The reason why resides in the size of gene-space. For our genome is length two, mutations cancelling each other out would be a fairly common occurence. Would cancelling out increase or decrease on a genome of length 1,000? Surely less. How much less (a forteriori!) the case for genomes with three billion molecules. By the extreme dimensionality of gene-space, then, we are witness to non-cancellative genetic movement!
- Genetic Drift Is (Random) Travel.
Importantly, it is not the individuals that travel (modify their genomes), but the species as a whole.
- Species Are Vehicles.
Viewing the species itself as actor, rather than the individual, is an important paradigm shift of population genetics.
Takeaways
In this post, I introduced the following metaphor:
- A Genotype Is A Location.
- Organisms Are Unmoving Points
- Birth Is Point Creation, Death Is Point Erasure
- Genome Differences Are Distances
We then strengthened our metaphor with the following considerations:
- A Species Is A Cluster Of Points
- Species Are Vehicles
- Genetic Drift is (Random) Travel.
We are left with the image of specie vehicles clumsily moving around gene-space. But genetic drift is not the only mechanism by which species navigate gene-space. In our next post, we explore a more sophisticated property of living things.